Clones, subclones, malignant clones, clonal architecture, clonal expansion, clonal progression, clonal selection, clonal evolution… the terminology can be confusing. They often create uncertainty for both patients and physicians who are trying to interpret laboratory results involving mutations, allele frequencies, and pathogenicity.
Once considered a largely theoretical principle, clonality is now central to contemporary hematology, oncology, immunology, and molecular diagnostics. This article explores the fundamentals of clonality, recognizing its potential for malignant evolution without overstating that risk, and how modern science is transforming its role in risk prediction and personalized medicine.
What Makes a Clone a Clone?
Clonality refers to the presence of a cell population derived from one common progenitor cell. These descendant cells inherit biological features from the founding cell, including genetic mutations, epigenetic patterns, or characteristic receptor rearrangements.
In practical terms, it means that one cell acquires characteristics that allow it to proliferate and generate a population of genetically related descendants. This principle underlies many malignancies, yet it is not limited to cancer. It also reflects normal immune responses, stem-cell competition, and the biology of aging:
1. When the immune system encounters an antigen, selected B lymphocytes and T lymphocytes proliferate rapidly. This creates antigen-specific clonal expansion, an essential mechanism for adaptive immunity.
2. Normal tissues maintained by stem cells may undergo gradual shifts in clonal contribution over time. Some stem-cell lineages become more productive than others, a phenomenon sometimes called clonal drift.
3. With aging, somatic mutations accumulate in long-lived stem cells. Some mutated cells gain a competitive advantage and expand, particularly within the hematopoietic system.
Several related terms are commonly used:
- Monoclonal – derived predominantly from one ancestral cell
- Polyclonal – derived from multiple independent cells
- Oligoclonal – composed of several dominant clones
This distinction is clinically important. Reactive or inflammatory processes are often polyclonal, whereas neoplastic disorders frequently show monoclonality. Thus, clonality is not inherently pathological, it is a process that can be physiological or disease-associated depending on context.
Why Darwinian?
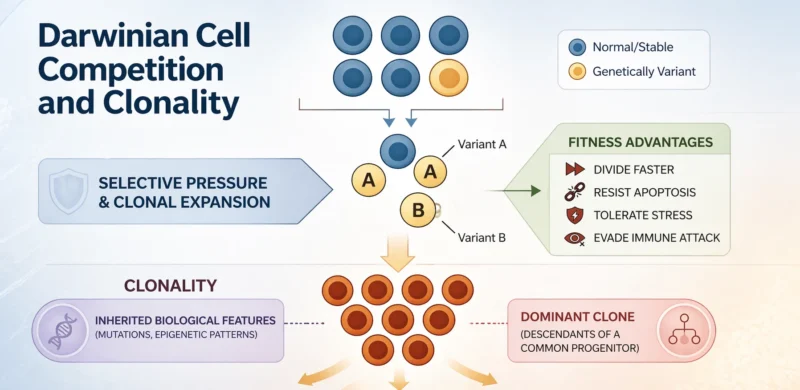
Darwinian Cell Competition refers to the idea that cells within the body can undergo a process analogous to Charles Darwin’s theory of natural selection: cells compete with one another, and those with traits that confer a survival or growth advantage become dominant over time.
Inside tissues (which might be bone marrow, blood, skin or tumors) many cells coexist. If one cell acquires a mutation that helps it:
- divide faster
- resist apoptosis
- better tolerate stress
- evade immune attack
- survive therapy
Clonality is a dynamic evolutionary process in which genetically distinct cell populations compete under selective pressure. This natural selection occurring at the cellular level creates clonal expansion.
Why Hematology Is Well Suited to Study Clonality?
Blood disorders have become a model system for translational clonality research because hematopoiesis is dynamic, hierarchical, and clinically accessible. Peripheral blood and bone marrow can be sampled repeatedly, allowing direct observation of clone behavior over time.
How Is Clonality Detected?
Historically, clonality was inferred through indirect methods such as morphology, cytogenetics, flow cytometry, or X-chromosome inactivation assays. In hematology, recurrent chromosomal abnormalities or a uniform immunophenotype often suggested that a population of cells was clonal rather than polyclonal. These earlier approaches remain useful, but they are less sensitive than modern molecular techniques.
Today, the most common method is NGS of peripheral blood, bone marrow, or tumor tissue. Targeted sequencing panels can detect recurrent somatic mutations in genes such as DNMT3A, TET2, ASXL1, TP53, JAK2, and spliceosome genes, allowing identification of clonal hematopoiesis or overt myeloid neoplasia. The proportion of cells carrying a mutation is often estimated using the variant allele frequency (VAF), which serves as a practical surrogate for clone size.
Cytogenetics and chromosomal microarrays detect gains, losses, or structural rearrangements, while single-cell DNA/RNA sequencing can reconstruct clonal architecture by showing which mutations coexist within the same cell and how subclones relate to one another.
More recently, cell-free DNA (cfDNA) analysis has emerged as a minimally invasive “liquid biopsy” approach. Plasma sequencing can detect circulating tumor-derived variants and some hematopoietic clones, although distinguishing tumor mutations from clonal hematopoiesis remains a challenge.
Current evidence suggests cfDNA is promising for monitoring disease dynamics, but matched blood-cell sequencing is often needed for accurate interpretation. In practice, clonality is therefore best assessed by integrating molecular results with blood counts, marrow morphology, clinical context and longitudinal follow-up.
Clonality vs Cancer: Not Every Clone Is Malignant
Approximately 50 genes are recurrently mutated in MDS and AML. Is the presence of such a mutation in an otherwise healthy individual preleukemia?
Clonal Hematopoiesis of Indeterminate Potential (CHIP), was coined to categorize this laboratory finding and distinguish it from true myeloid neoplasia. Most individuals with CHIP never progress, and many hematologic cancers still arise without previously detectable CHIP.
In a risk-stratification study, Lachelle D. Weeks noted that the absolute risk of myeloid malignancy in CHIP remains about 0.5-1% per year, despite a higher relative risk than the general population. Progression risk was concentrated in subsets with larger clone size, multiple mutations, cytopenias, or high-risk genes such as spliceosome mutations, TP53, IDH1/2, FLT3 and RUNX1, whereas isolated DNMT3A-mutant CHIP carried substantially lower risk.
In MGUS, a single plasma-cell clone produces a measurable monoclonal immunoglobulin (M-protein), yet patients lack the defining features of multiple myeloma. Recent reviews continue to show that MGUS is common with aging and carries an average progression risk of roughly 1% per year, meaning that most individuals never develop overt plasma-cell cancer.
MBL is characterized by a small clonal B-cell population in the peripheral blood without the diagnostic clinical features of CLL or lymphoma. It is frequently discovered incidentally, especially in older adults and many low-count cases remain stable for years without treatment. Higher-count MBL carries a greater, but still limited, risk of progression (Zanwar S et al., 2025).

Further evidence comes from studies of bone marrow failure syndromes. Paroxysmal Nocturnal Hemoglobinuria provides a classic example of benign clonal hematopoiesis. It arises from acquired PIGA mutations, and the clone can be readily detected by flow cytometry. Small PIGA-mutated stem-cell clones may even be found in otherwise healthy individuals, yet progression to overt PNH remains uncommon.
A similar complexity is seen in acquired Aplastic Anemia, where many patients respond to immunosuppressive therapy, but a minority later develop MDS or AML. Some studies have suggested that detectable somatic mutations predict later clonal evolution, particularly mutations in ASXL1.
Long-term follow-up further shows that somatic clones in AA may expand, shrink, or remain stable over years, often with little correlation to blood counts or immediate clinical change. Some clones with higher-risk mutations remain small and indolent, whereas others evolve unpredictably. Larger collaborative analyses found that the mere presence of a mutation was not an independent predictor of progression. Instead, risk appeared to depend more on the specific mutational pattern than on clonality alone (Cooper J.N. et al., 2017).
When Clonality Becomes Malignant
Cancer commonly begins when a progenitor cell acquires one or more driver mutations that confer a selective advantage in growth, survival, self-renewal, or resistance to normal regulatory signals. Once such a change occurs, the altered cell can expand disproportionately relative to neighboring cells, giving rise to a founding clone.
However, malignant clonality is rarely static. As the founding clone proliferates, descendant cells continue to acquire additional genetic lesions. This generates subclonal diversity, with multiple related but genetically distinct populations coexisting within the same disease.
This evolutionary process mirrors Darwinian selection: clones compete within a constrained microenvironment, and those with the greatest fitness dominate over time. Treatment itself can become a selective pressure, eliminating sensitive populations while allowing resistant subclones to expand. Clinically, this helps explain relapse, progression, and transformation after an initial response.
Modern sequencing studies therefore suggest that cancer should not be viewed as a single uniform clone, but rather as a dynamic ecosystem of competing cellular lineages undergoing continuous evolution. Understanding this clonal architecture has become central to prognosis and precision strategies.
In a systematic review and meta-analysis by Kowalska M et al. involving 32 studies and 56 cohorts, detectable CH mutations were associated with a higher risk of hematologic cancer compared with individuals without CH (HR 5.59, 95% CI 3.31-9.45). Across the analyzed studies, malignant risk increased stepwise with higher VAF thresholds. Smaller clones detected at low VAF (≥0.5%) were associated with modest risk elevation, whereas larger clones (≥10% VAF) showed markedly higher risk, with hazard ratios reaching 49 in some cohorts.
Clonal Hematopoiesis in Cancer Cohorts: Risk Factors and Germline Drivers
CH appears to be more common in cancer cohorts, likely reflecting prior exposure to mutagenic stressors such as chemotherapy, radiation or transplant conditioning. Cytotoxic therapy imposes strong selective pressure on the hematopoietic stem-cell compartment, eliminating many normal progenitors while favoring survival of clones with mutations that confer resistance to DNA damage or apoptosis. As a result, hematopoietic recovery after treatment may become increasingly driven by these therapy-selected clones rather than by a fully polyclonal stem-cell pool.
Genomic studies have shown that many therapy-related clones were already present at very low levels before cancer treatment and subsequently expanded after exposure to cytotoxic stress. These clones are enriched for mutations in TP53, PPM1D, and other genes involved in DNA-damage response. Clinically, therapy-related clonality is important because it is linked to a higher risk of subsequent myeloid neoplasms, inferior survival, and abnormal hematopoietic recovery after treatment. These findings have challenged the traditional view that therapy-related MDS/AML arises solely from new treatment-induced mutations.
Additional risk factors, in both individuals with and without cancer, include current or former smoking and possible environmental mutagen exposure. Population-based genomic studies have shown that clonal hematopoiesis is enriched in individuals carrying inherited variants near TERT, the gene encoding telomerase. People with JAK2 V617F-mutant clonal hematopoiesis, have identified predisposition variants near TERT, JAK2, and to a lesser extent TET2. Familial clustering of TET2-mutant clonal hematopoiesis further supports a hereditary component.
How Should Incidental CH Be Managed in Cancer Patients?
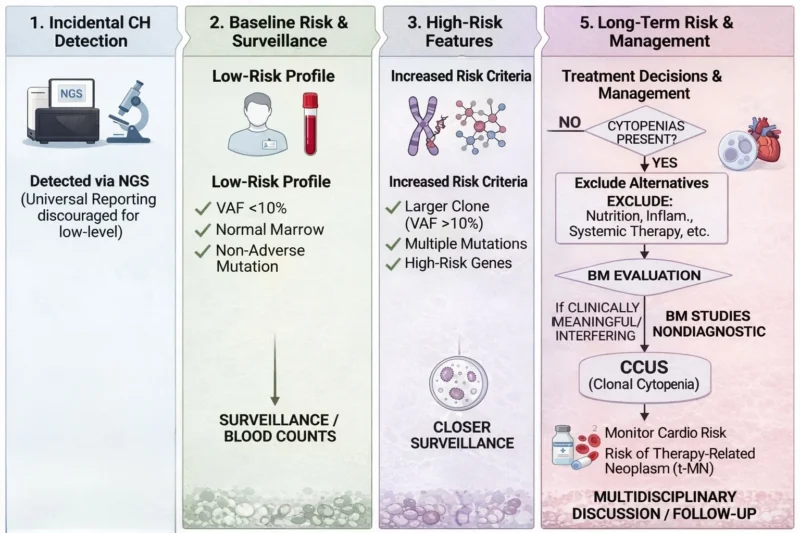
The incidental detection of CH in patients undergoing molecular testing for solid tumors or lymphoma is becoming common with the broader use of NGS. Current expert perspectives do not support universal reporting of every low-level CH finding, particularly when marrow function is normal and the clone is limited to a single mutation in a non-adverse gene at VAF <10%. Interpretation should always occur within the broader clinical context.
When abnormal blood counts are present, physicians should first exclude more common alternative causes such as nutritional deficiency, inflammation, infection, marrow suppression from systemic therapy, or organ dysfunction. If cytopenias are clinically meaningful or interfere with oncologic treatment, bone marrow evaluation is appropriate.
When no alternative explanation is found and marrow studies remain nondiagnostic, the patient may meet criteria for clonal cytopenia of uncertain significance (CCUS), a state believed to carry higher risk of progression to myeloid neoplasia than asymptomatic CH alone. Conversely, patients without cytopenias but with larger clones (VAF >10%), multiple mutations, or lesions involving higher-risk genes may need a closer hematologic surveillance.
More complex decisions arise when patients with CH are expected to receive additional cytotoxic therapy, stem-cell transplantation, or other selective pressure. Management should be individualized, balancing the urgency of cancer treatment against future hematologic risk. Follow-up blood counts, multidisciplinary discussion and attention to associated cardiovascular risk factors are also prudent, as CH increasingly appears to have implications beyond hematologic malignancy alone (Park S.J. et al., 2020, J Mol Pathol, 2025).
A Window for Prevention
Inflammation is increasingly recognized as a major driver of clonal expansion, particularly in therapy-related CH. Chronic inflammatory stress may selectively favor mutant hematopoietic stem cells, which often display greater resistance to suppressive cytokines. These clones can further reshape the marrow niche through pro-inflammatory signaling, reinforcing their own growth. Together, these findings raise the possibility that CH may represent a modifiable risk state, in which restoring a healthier microenvironment could help limit further clonal evolution.
One of the best-characterized examples involves TET2-mutant CH, which has been linked to overproduction of IL-1β and atherogenic inflammation. In the CANTOS trial, the IL-1β inhibitor canakinumab produced only modest overall cardiovascular benefit, but subsequent analyses suggested much of that effect occurred in patients with TET2-mutant CH.
In Tet2-knockout mouse models, inhibition of inflammatory signaling reduced the survival advantage of mutant clones, while antibiotic-mediated microbiome modulation also lowered Tet2-mutant clonal burden. These observations have prompted interest in accessible interventions such as metformin and statins, as well as more specific targets including the NLRP3 inflammasome, CD33, and S100A9, all implicated in myeloid disease biology.
Molecularly targeted prevention strategies are also being considered. IDH1/2-mutant CH carries relatively high risk of progression to AML, raising theoretical interest in earlier use of IDH inhibitors. Similarly, TP53-mutant CH has prompted discussion of agents such as eprenetapopt (Park S.J. et al., 2020).
Next Frontiers in Clonality Research
One of the most important advances is the rise of single-cell multi-omics, which allows simultaneous analysis of genomic, transcriptomic, epigenetic, and proteomic features within individual cells rather than bulk populations. This distinction is critical, as conventional bulk sequencing may identify the presence of mutations but often obscures how they are distributed across cellular compartments and developmental states.
Recent studies in hematopoiesis and AML have demonstrated how single-cell platforms can uncover previously hidden evolutionary trajectories, including rare preleukemic stem-cell populations and treatment-resistant residual clones.
A parallel frontier is the use of artificial intelligence and computational evolutionary modeling. As sequencing datasets become larger and more complex, machine-learning approaches are increasingly needed to infer clonal fitness, predict progression risk, and simulate future evolutionary paths under therapeutic pressure. Rather than describing disease retrospectively, these tools may help forecast which clones are most likely to expand, relapse, or acquire resistance.
Finally, clonality is likely to become central to personalized treatment selection, with therapies tailored according to clonal architecture and subclonal diversity. Integrating aging biology into hematologic risk prediction may further refine care, particularly as age-related clonal expansions become more common in older populations (Beneyto-Calabuig S. et al., 2023, Liu J. et al., 2024, Scherer M. et al., 2025)
Together, these advances suggest that future medicine may treat not only diseases themselves, but also the clonal ecosystems from which they arise.
You can also read: Myelodysplastic Neoplasms (MDS): From Ineffective Hematopoiesis to Sideroblastic Phenotype
Written by Susanna Mikayelyan, MD
FAQ
What is clonality?
Clonality refers to the presence of a population of cells derived from one common ancestral cell. These related cells often share genetic or biological features inherited from the founding clone.
What is the difference between clonality and cancer?
Clonality means a group of cells arose from one ancestral cell, but that does not automatically mean cancer. Many clonal states are benign, age-related, or remain stable for years without progression.
Can a healthy person have clonal hematopoiesis?
Yes. Clonal hematopoiesis is increasingly detected in otherwise healthy individuals, especially with aging. Most people with low-risk clones never develop a blood cancer.
Why does clone size matter in hematology?
Clone size, often estimated by variant allele frequency (VAF), can provide insight into biological significance. Larger clones are generally associated with higher risk than very small incidental clones.
Why are blood diseases ideal for studying clonality?
Blood and bone marrow can be sampled repeatedly over time, allowing researchers to directly observe how clones expand, regress, or evolve under treatment pressure.
What is clonal evolution?
Clonal evolution refers to the process by which cell populations acquire additional mutations over time, leading to new subclones that may become more aggressive or treatment-resistant.
Can cancer treatment unintentionally select harmful clones?
Yes. Chemotherapy, radiation, and other cytotoxic therapies can eliminate sensitive cells while allowing resistant pre-existing clones to expand, especially those with TP53 or PPM1D mutations.
Could clonality become a target for prevention?
Possibly. Researchers are studying whether high-risk clones can be monitored or intercepted before overt malignancy develops using anti-inflammatory, metabolic, or molecularly targeted strategies.