Cancer is increasingly recognized not as a collection of malignant cells, but as a complex and spatially organized ecosystem. Within tumors, malignant cells coexist with immune populations, stromal components, and extracellular matrix elements, forming a dynamic network of interactions.
These interactions are not random. Their spatial organization directly influences tumor progression, immune evasion, and response to therapy. Traditional molecular approaches have provided critical insights into tumor biology, but they have been limited in their ability to capture this spatial dimension. Spatial omics has emerged to address this gap, enabling the study of gene and protein expression within intact tissue architecture
Technological Foundations of Spatial Omics
Spatial omics encompasses a range of technologies that preserve tissue structure while profiling molecular information.
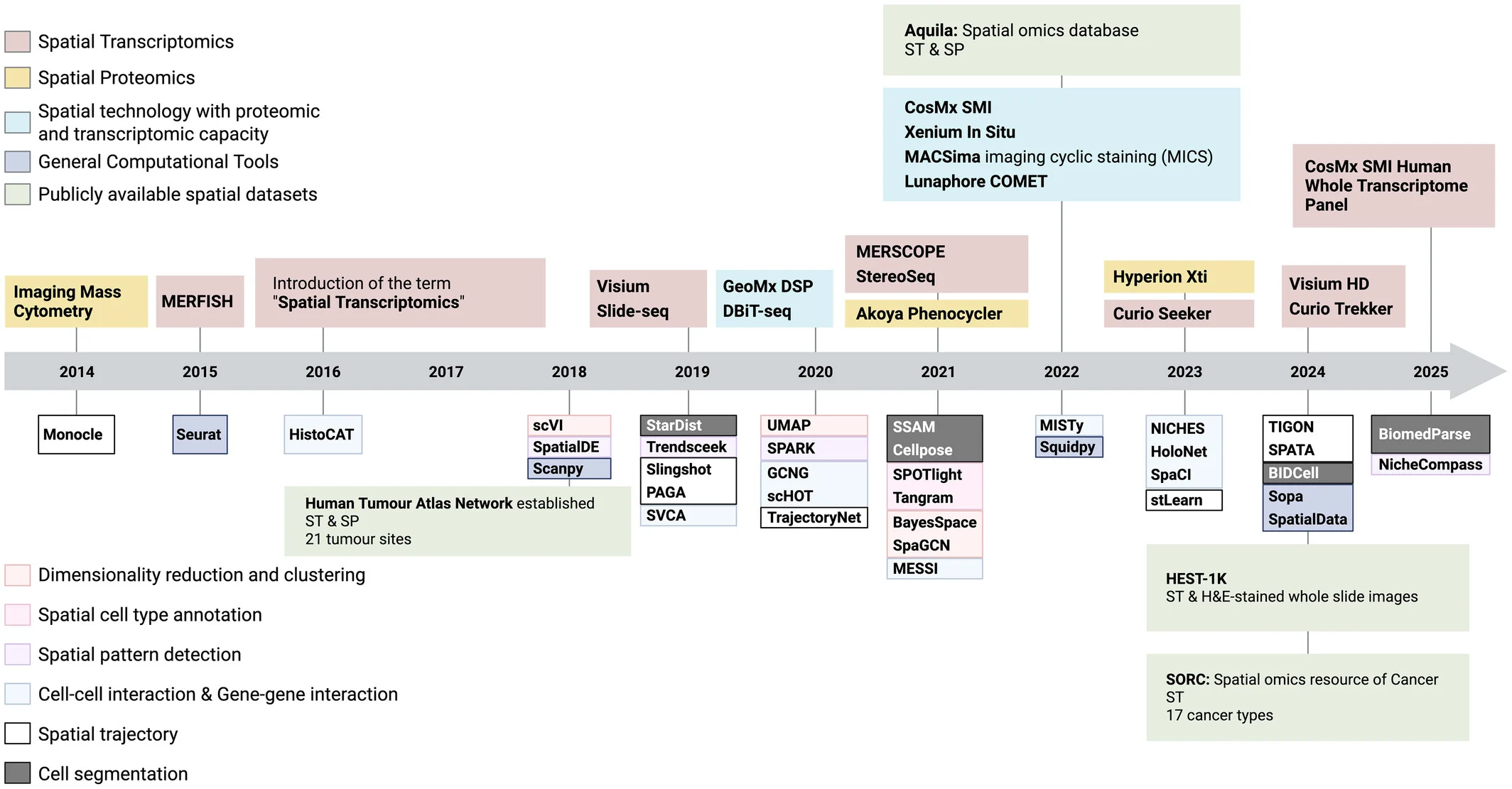
Spatial transcriptomics technologies can be broadly divided into sequencing-based and imaging-based approaches. Sequencing-based platforms enable genome-wide transcriptomic profiling across tissue sections, while imaging-based methods provide single-cell and even subcellular resolution. In parallel, spatial proteomics technologies allow mapping of protein expression within tissue context.
Together, these approaches create a multi-dimensional view of tumors, integrating structural organization, transcriptional activity, and functional protein expression within the same spatial framework.
From Molecular Data to Spatial Biology
The key advancement of spatial omics lies in its ability to move beyond molecular description toward spatial understanding.
Bulk sequencing approaches average signals across heterogeneous tissues, masking regional variability. Single-cell technologies improve resolution but disrupt tissue architecture, removing information about cellular neighborhoods. Spatial omics bridges this gap by preserving both resolution and context.
This enables the reconstruction of cellular neighborhoods, identification of spatial domains, and mapping of cell–cell interactions within the tumor microenvironment. As a result, tumor biology can now be studied as a system of interacting components rather than isolated molecular events.
Spatial Organization of the Tumor Microenvironment
Spatial omics has revealed that the tumor microenvironment is highly structured and heterogeneous.
Within a single tumor, multiple distinct regions may coexist, including immune-infiltrated areas, immune-excluded zones, fibroblast-rich barriers, and metabolically specialized niches. Cells behave differently depending on their spatial location, and these differences play a critical role in shaping tumor behavior.
This spatial heterogeneity helps explain variability in treatment response, even within the same tumor, and highlights the importance of local cellular context in determining disease outcomes.
Spatial Dynamics of Tumor Progression
Tumor progression is increasingly understood as a spatially coordinated process.
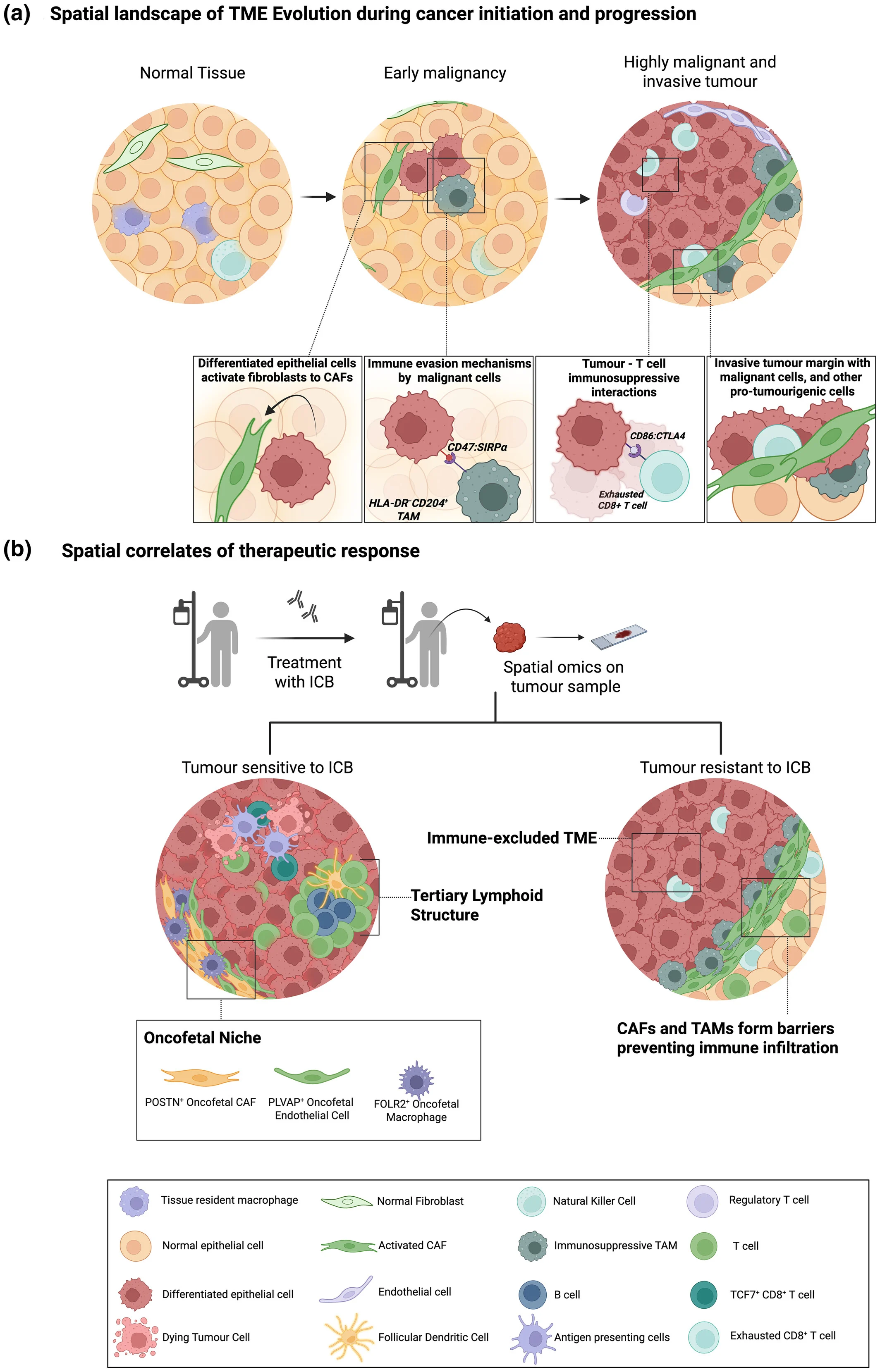
Early transformation often occurs within localized niches, where interactions between epithelial cells, immune populations, and fibroblasts drive tumor initiation. As tumors evolve, distinct spatial regions emerge with specialized biological functions, including invasive fronts and immunosuppressive microenvironments.
Spatial omics studies have demonstrated that these region-specific interactions are central to cancer progression, indicating that tumor evolution is shaped not only by genetic alterations but also by its surrounding microenvironment.
Implications for Immunotherapy
One of the most clinically significant contributions of spatial omics is in immuno-oncology.
Response to immunotherapy depends not only on the presence of immune cells, but on their spatial organization and proximity to tumor cells. Tumors that respond to immune checkpoint blockade often exhibit structured immune niches and direct tumor–immune interactions. In contrast, resistant tumors frequently display spatial separation, where immune cells are excluded or confined to stromal compartments.
Spatial features such as tertiary lymphoid structures and organized immune cell clusters are emerging as important biomarkers of response. This represents a shift from measuring immune cell quantity to understanding immune architecture.
Implications for Immunotherapy
One of the most clinically relevant contributions of spatial omics lies in immuno-oncology.
Response to immunotherapy depends not only on the presence of immune cells, but on their spatial organization and proximity to tumor cells. Tumors that respond often exhibit coordinated immune structures and direct tumor–immune interactions, whereas resistant tumors frequently show spatial separation.
This shift reframes biomarker discovery—from measuring immune quantity to understanding immune architecture.
Computational and AI Integration
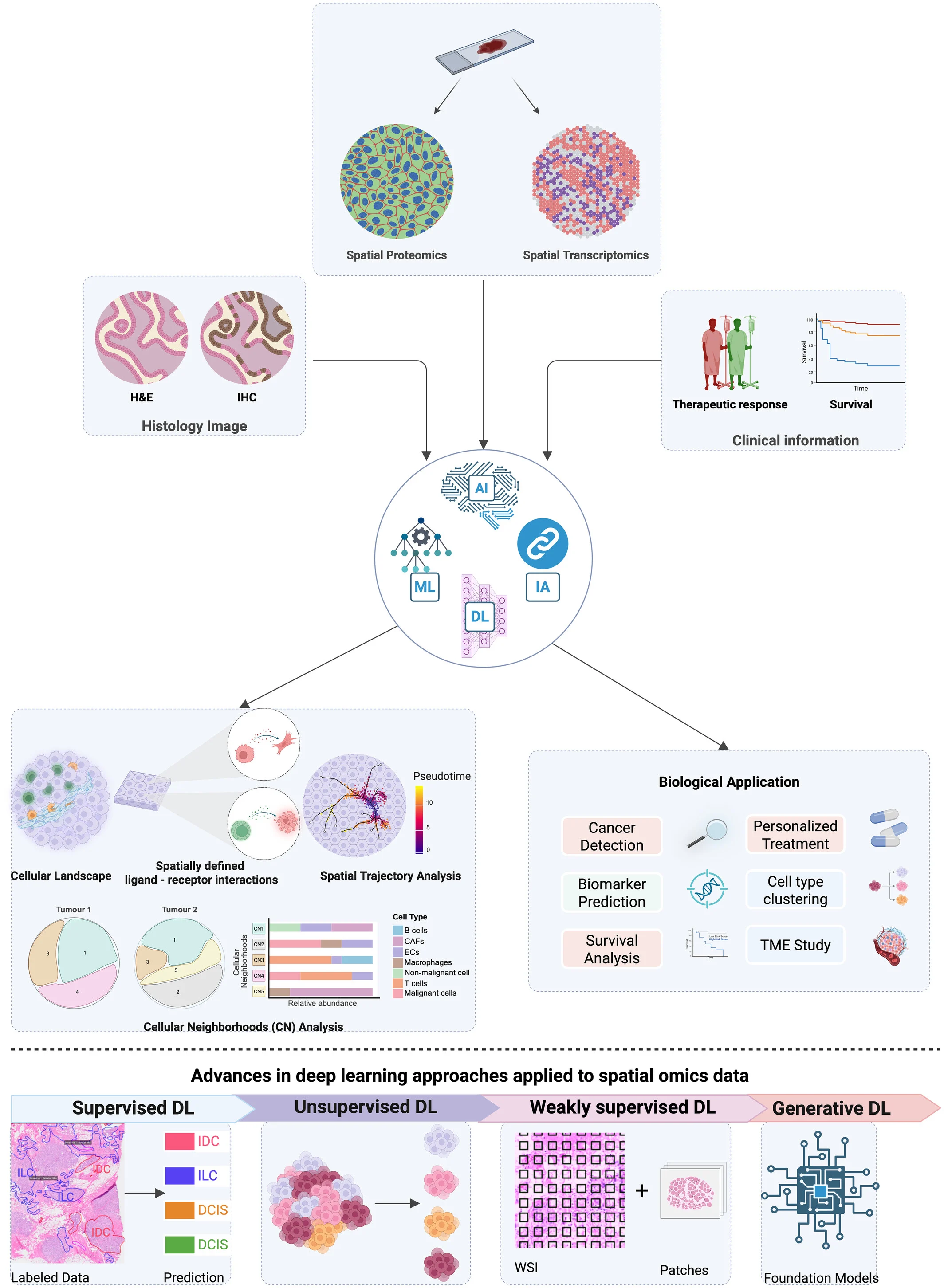
The complexity of spatial omics data requires advanced computational approaches for meaningful interpretation. Analytical pipelines enable normalization, clustering, spatial domain identification, and inference of cell–cell communication networks. Increasingly, artificial intelligence and machine learning are used to integrate spatial data with histopathology and clinical outcomes, enabling prediction of treatment response and identification of novel biomarkers.
Despite these advances, challenges remain in standardization, scalability, and interpretability, which are essential for clinical translation.
Challenges and Barriers to Clinical Implementation
Although spatial omics holds significant promise, its integration into routine clinical practice remains limited.
Key barriers include high cost, technical complexity, lack of standardized workflows, and difficulties in integrating data across platforms. Additionally, prospective validation in clinical trials is required to establish the clinical utility of spatial biomarkers.
Addressing these challenges will be critical for translating spatial omics from research into clinical oncology.
Future Directions: Toward Spatial Precision Oncology
Spatial omics is rapidly transitioning from discovery research to translational application.
Future developments are expected to focus on identifying clinically actionable spatial biomarkers, integrating spatial data into pathology workflows, and combining spatial omics with other molecular and clinical data. These advances will support more precise and context-aware treatment strategies.
Conclusion
Spatial omics represents a conceptual shift in cancer research. Tumors are no longer viewed solely as genetically defined entities, but as spatially organized and evolving ecosystems.
Understanding cancer now requires not only sequencing its molecular components, but also mapping their spatial distribution and interactions within tissue. This approach provides a more comprehensive view of tumor biology and has the potential to transform precision oncology.
In this evolving paradigm, biology is defined not only by what is present—but by where it exists and how it interacts.
You Can Read Full Article Here