Marios Georgakis, Consultant Neurologist at LMU Klinikum München, Research Group Leader at Ludwig-Maximilians-Universität München and Principal Investigator at Deep Vascular Phenotyping (DeepVasc) Lab, shared a post on LinkedIn:
“While the genetic revolution promised to return thousands of drug targets, after massive investments in genomic and multi-omic datasets, we still struggle interpreting individual signals.
These recent comments in Nature Genetics on a GWAS signal for cerebral amyloid angiopathy at APOE are a great case study in the challenges of interpreting genetic associations even with abundant data. Worth reading for anyone interested in translating GWAS hits to drug targets.
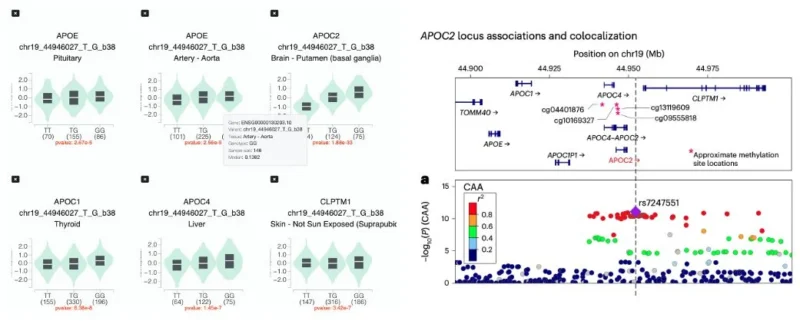
The lead risk variant (rs7247551) is highly correlated with a variant (rs2288911) that increases APOE expression in microglia and is associated with both Alzheimer’s disease and cerebral amyloid angiopathy. Increased microglial APOE expression as a mechanistic explanation aligns well with experimental studies and might be biologically appealing.
However, the same variant also affects the expression of other neighboring genes (APOC1, APOC2, APOC4, CLPTM1) across several tissues, including brain regions. Interestingly, for APOC2, the expression effect in brain appears stronger than for APOE. Supporting this, the variant is associated with altered DNA methylation at CpG sites upstream of APOC2 in dorsolateral prefrontal cortex, and the signal colocalizes with APOC2 expression across brain regions.
Taken together, despite the availability of large-scale GWASs for clinical phenotypes and neuropathology, extensive transcriptomic resources across tissues and brain regions, and even single-cell eQTL data, we still cannot draw a clear conclusion about the causal gene or mechanism at this locus.
If interested in pharmacologically exploiting the driving signal, this matters, as targeting microglial APOE versus APOC2-driven pathways would imply fundamentally different drug development strategies.
A good reminder that even in the era of massive datasets and rich functional data moving from genetic association to mechanism, and ultimately to actionable drug targets, remains more challenging than discovering new loci.”
Title: A quantitative trait locus for reduced microglial APOE expression associates with reduced cerebral amyloid angiopathy
Authors: Michael E. Belloy, Jonathan Graff-Radford, Michael D. Greicius
Read the Full Article.

Title: Reply to: a quantitative trait locus for reduced microglial APOE expression associates with reduced cerebral amyloid angiopathy
Authors: Lincoln M. P. Shade, Qi Qiao, Yuriko Katsumata, Shubhabrata Mukherjee, Jai G. Broome, Lance A. Johnson, Mark T. W. Ebbert, Peter T. Nelson, David W. Fardo
Read the Full Article.

Other articles featuring Marios Georgakis on OncoDaily.


