Kinase inhibitors have transformed the treatment of oncogene-driven lung cancer, from EGFR-mutated non-small cell lung cancer to ALK-, RET-, ROS1-, BRAF-, MET-, and NTRK-altered disease. Yet one of the persistent challenges in thoracic oncology is that kinase biology is rarely static. Tumors acquire resistance mutations, different variants respond differently to available inhibitors, and clinically approved kinase drugs often inhibit targets beyond the ones for which they were originally developed. A new Nature Biotechnology resource by Saifudeen and colleagues provides one of the most comprehensive functional maps of this problem, profiling clinically approved kinase inhibitors across hundreds of wild-type and oncogenic kinase variants and creating a public tool, KIRHub, to support precision oncology and drug repurposing (Saifudeen et al., 2026).
For lung cancer, the work is especially relevant. Lung tumors represented the largest mutation burden among the profiled cancer-associated kinase variants, and the study directly examined several of the most important lung cancer kinase families, including EGFR, ALK, RET, MET, ROS1, BRAF, BTK, and KIT (Saifudeen et al., 2026). The findings do not immediately change clinical practice, but they provide a powerful research framework for understanding mutation-specific inhibitor sensitivity, resistance gaps, and potential repurposing opportunities in kinase-driven lung cancer.
Why Kinase Profiling Matters In Lung Cancer
Protein kinases are central regulators of cell signaling, proliferation, differentiation, metabolism, migration, and apoptosis. In cancer, dysregulated kinase activity can support several hallmarks of malignancy, including uncontrolled growth, survival signaling, invasion, metastasis, and immune suppression (Saifudeen et al., 2026). This is why kinase inhibitors have become a major therapeutic class in oncology.
In lung cancer, kinase targeting is already part of routine precision oncology. EGFR inhibitors, ALK inhibitors, RET inhibitors, MET inhibitors, ROS1 inhibitors, and BRAF-directed strategies have created biomarker-defined treatment pathways. However, the effectiveness of these therapies is often shaped by the exact molecular alteration. A drug that inhibits one variant of a kinase may not inhibit another. Secondary resistance mutations can also emerge under treatment pressure, requiring a different inhibitor strategy.
The study by Saifudeen et al. addresses this problem by moving beyond single-drug, single-target thinking. Instead, it asks a broader question: among clinically approved kinase inhibitors, which drugs inhibit which wild-type kinases, which oncogenic mutations, and which kinase fusions?
A Large Functional Map Of Approved Kinase Inhibitors
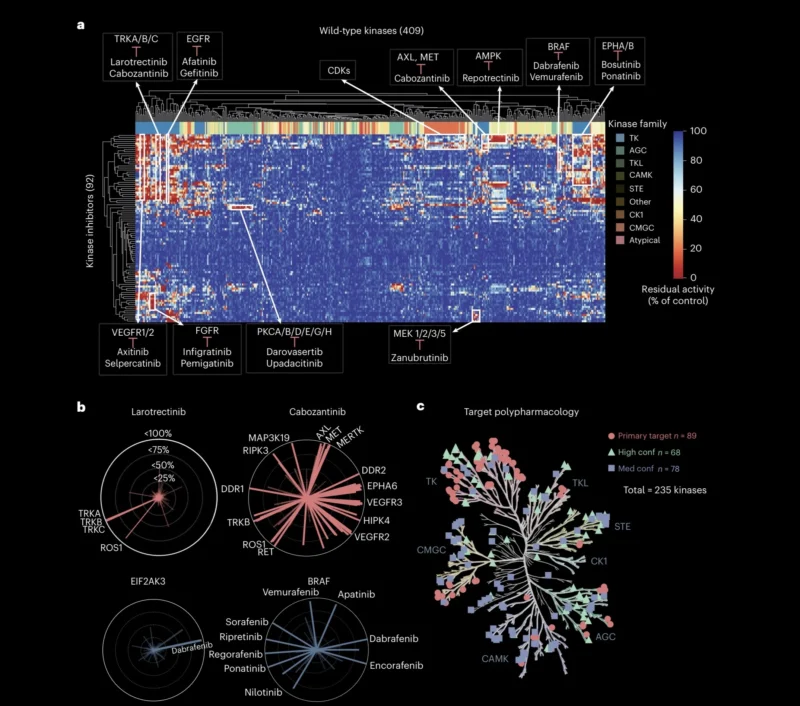
The investigators profiled 92 clinical kinase inhibitors across 758 kinases, including 409 wild-type kinases and 349 mutant kinases or kinase fusion products. Of the 92 inhibitors, 86 were FDA-approved, one was approved by China’s National Medical Products Administration, and five were in late-stage clinical trials (Saifudeen et al., 2026).
The work used the HotSpot radiometric filter-binding kinase assay, a biochemical platform that directly measures kinase-mediated phosphate transfer from ATP to a protein or peptide substrate. The authors selected 1 μM as the screening concentration after dose-response modeling, allowing them to capture both primary and secondary kinase targets while maintaining meaningful selectivity (Saifudeen et al., 2026).
In total, the study generated approximately 290,000 kinase–drug interaction measurements, creating what the authors describe as the largest functional kinase inhibitor dataset to date (Saifudeen et al., 2026). Importantly, the dataset was not limited to wild-type kinases. It included oncogenic mutations and fusion variants, which is critical for lung cancer because resistance and sensitivity are often mutation-specific.
Expanding The Druggable Kinase Landscape
One of the most important findings was that approved kinase inhibitors have a much broader target landscape than their labeled targets suggest. The 92 clinical kinase inhibitors were originally designed to target 89 primary kinases, but the profiling analysis expanded the druggable kinase landscape to 235 kinases when secondary targets were included (Saifudeen et al., 2026).
This reflects kinase inhibitor polypharmacology, the ability of one drug to inhibit multiple kinases. Traditionally, polypharmacology has often been viewed as a liability because off-target activity can contribute to toxicity. However, this study emphasizes a more nuanced view: when understood and mapped systematically, polypharmacology may create opportunities for drug repurposing, pathway-level inhibition, or overcoming resistance.
For lung cancer, this concept is particularly important because resistance frequently occurs through kinase-domain mutations, pathway bypass, or altered target dependence. A drug developed for one kinase family may unexpectedly inhibit another clinically relevant mutant kinase, creating hypotheses for future testing.
Lung Cancer Had The Largest Kinase Variant Representation
The study profiled 349 kinase variants, including 311 mutations and 38 gene fusions. These variants spanned multiple tumor types, but lung cancer contributed the highest number of unique kinase mutations, with more than 150 represented in the dataset (Saifudeen et al., 2026).
This makes the resource especially useful for thoracic oncology. Lung cancer is one of the most genotype-driven solid tumors, and treatment decisions increasingly depend on the precise mutation or fusion present. The ability to examine whether an approved inhibitor biochemically inhibits a specific EGFR, ALK, RET, MET, ROS1, BRAF, BTK, or KIT variant can help generate rational hypotheses for translational research, resistance studies, and future trial design.
The authors found that 94% of kinase mutations and 97% of kinase fusions in their profiled set were inhibited by at least one existing drug with more than 70% inhibition activity (Saifudeen et al., 2026). This does not mean that all such variants are clinically targetable today, but it suggests that many oncogenic kinase variants may have potential inhibitor matches among existing drugs.
EGFR, ALK, And RET Show Variant-Specific Patterns
The authors examined three of the most commonly altered kinase families in lung cancer: ALK, EGFR, and RET. They assessed approved inhibitors against 21 ALK mutants or fusion products, 64 EGFR mutants, and 33 RET mutants or fusion products, using a threshold of less than 20% residual activity to define effective inhibition (Saifudeen et al., 2026).
For ALK, approved inhibitors such as crizotinib, ceritinib, brigatinib, alectinib, and lorlatinib showed activity across several ALK variants and fusion products. For EGFR, inhibitors including lazertinib, osimertinib, gefitinib, dacomitinib, erlotinib, and afatinib showed variable activity depending on the specific EGFR mutant. Some EGFR variants were inhibited by multiple drugs, while others were inhibited by fewer agents or only one (Saifudeen et al., 2026).
This finding aligns with clinical reality in EGFR-mutated NSCLC: the exact EGFR alteration matters. Classical sensitizing mutations, exon 20 insertions, T790M, C797S-containing compound mutations, and other uncommon variants do not behave identically. The study reinforces that EGFR should not be treated as a single target category when resistance and uncommon mutations are being evaluated.
For RET, some mutations and most fusion products were inhibited by approved RET inhibitors, including selpercatinib, pralsetinib, and cabozantinib. The authors noted that in their analysis of ALK, RET, and EGFR, they did not identify a variant that was completely resistant to all currently approved drugs tested, although sensitivity varied substantially by variant (Saifudeen et al., 2026).
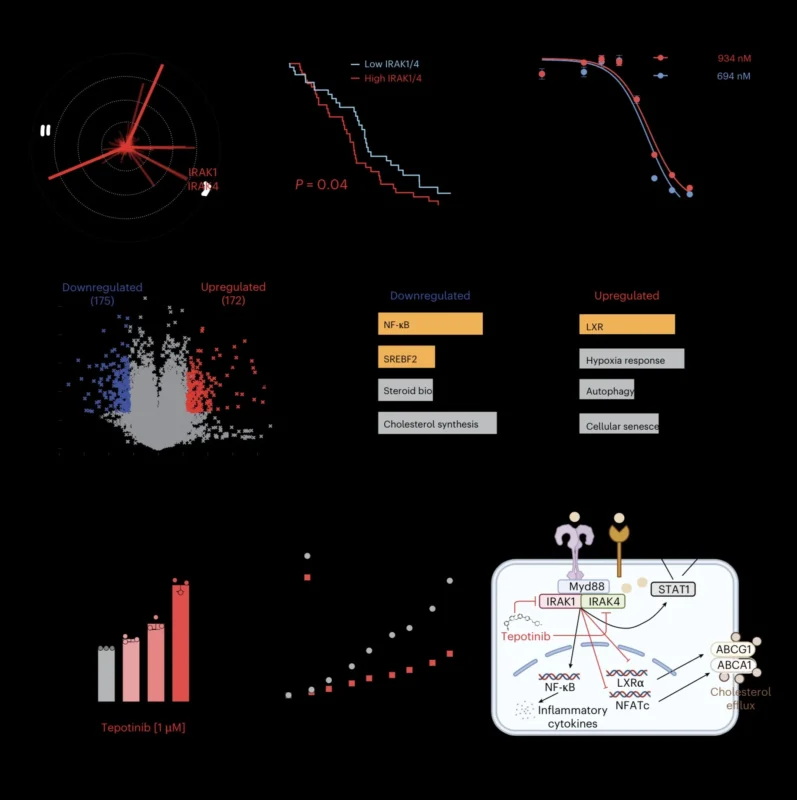
MET Resistance Mutations Reveal A Repurposing Opportunity
The MET findings may be among the most interesting for lung cancer drug resistance. MET exon 14 skipping and MET amplification are established therapeutic targets in NSCLC, and MET resistance mutations are increasingly relevant after MET inhibitor exposure.
The authors examined approved MET inhibitors, including crizotinib, tepotinib, capmatinib, and cabozantinib. They found that several MET variants, particularly mutations at D1228, were not effectively inhibited by approved MET inhibitors in the biochemical profiling dataset (Saifudeen et al., 2026). MET D1228 mutations are known resistance hotspots in MET-driven lung cancer.
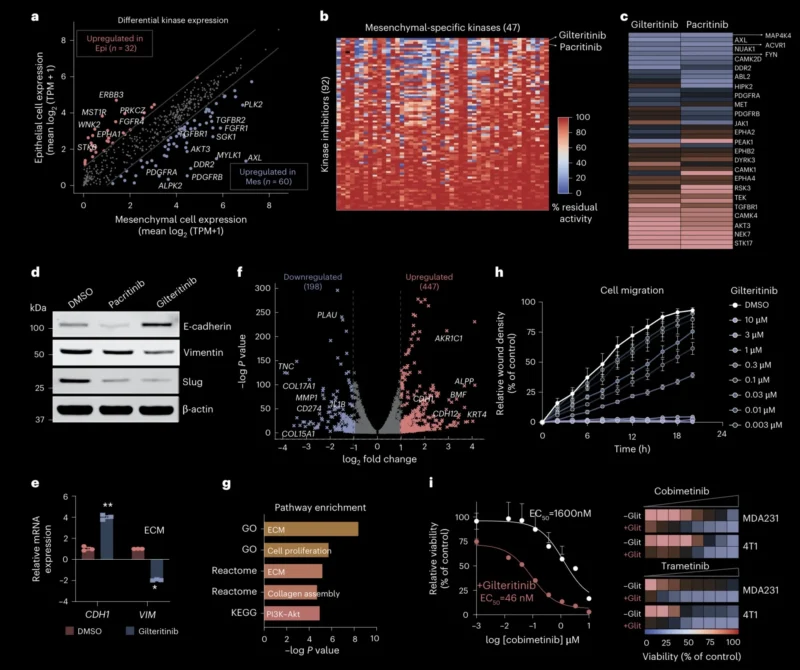
However, the study identified gilteritinib, an FLT3 inhibitor approved for FLT3-mutated acute myeloid leukemia, as active against multiple MET variants. In validation experiments, gilteritinib showed strong inhibition of MET-D1228A-expressing cells, with an EC50 of 0.21 μM, outperforming several approved MET inhibitors in that model (Saifudeen et al., 2026).
This is not a clinical recommendation to use gilteritinib for MET-mutated lung cancer. It is a preclinical and biochemical observation that requires cellular, translational, and clinical validation. Still, it is a valuable example of how systematic kinase profiling can identify potential repurposing candidates for resistant lung cancer variants.
FGFR Mutations Also Show Coverage Gaps
FGFR alterations are less common than EGFR or ALK alterations in lung cancer but remain relevant across solid tumors and occasionally in thoracic oncology. The study found that some FGFR variants were resistant to clinically approved FGFR inhibitors such as futibatinib, pemigatinib, and infigratinib (Saifudeen et al., 2026).
To address these gaps, the authors identified other clinically approved inhibitors with activity against resistant FGFR variants, including pralsetinib, selpercatinib, and erdafitinib. Selpercatinib target engagement with FGFR3-V555M was confirmed by cellular thermal shift assay, and additional validation supported the potential of these inhibitors against selected FGFR variants (Saifudeen et al., 2026).
Again, these findings are not immediately practice-changing, but they highlight how the “approved target” of a drug may not fully capture its biochemical potential. For precision oncology, that matters because rare variants often lack dedicated clinical trials.
KIRHub: A Public Tool For Precision Oncology Research
To make the dataset accessible, the authors developed KIRHub, the Kinase Inhibitor Repurposing Hub. This web-based tool allows users to explore inhibitors for mutant kinases, inhibitors for wild-type kinases, and kinase essentiality across cancer lineages (Saifudeen et al., 2026).
For lung cancer researchers, this type of tool may be useful for several purposes. It can help identify candidate inhibitors for rare or resistant kinase variants, compare drug activity across related kinase mutants, explore potential polypharmacology, and support translational hypotheses. The platform also allows results to be searched, sorted, visualized, and downloaded.
The key strength of KIRHub is that it provides a standardized biochemical reference map. The key caution is that biochemical inhibition does not automatically translate into clinical activity. Drug exposure, tumor context, cell permeability, toxicity, target expression, feedback signaling, and resistance biology all influence whether a biochemical hit becomes a useful treatment strategy.
Why This Resource Matters For Thoracic Oncology
The main lung cancer implication is not that every kinase variant now has a ready-to-use treatment. Rather, this study provides a structured way to ask better questions.
For uncommon EGFR mutations, the resource may help compare inhibitor sensitivity patterns. For ALK or RET fusions and resistance mutations, it may help identify whether current inhibitors retain activity across variants. For MET resistance, it provides a rationale for testing non-MET-labeled inhibitors such as gilteritinib in specific resistant contexts. For rare FGFR alterations, it suggests that inhibitors outside the expected target class may have activity against selected variants.
This is particularly relevant as lung cancer sequencing becomes more comprehensive. Next-generation sequencing increasingly identifies uncommon kinase mutations, compound resistance mutations, and rare fusions. Clinicians often face a challenge when a report identifies an alteration for which there is no standard treatment recommendation. Resources like KIRHub cannot replace clinical evidence, but they can support rational preclinical prioritization and trial development.
Important Limitations
The study has several limitations that are important for clinical interpretation. The profiling was performed using purified kinase constructs under standardized biochemical assay conditions. This allows cross-kinase comparison, but it does not capture tumor-cell context, kinase expression level, regulatory interactions, subcellular localization, drug metabolism, immune effects, or tumor microenvironment biology (Saifudeen et al., 2026).
The screening concentration of 1 μM was selected as a standardized biochemical reference, not as a direct model of clinical exposure. Lipid kinases such as PI3K were not included because of assay constraints. The mutation panel included 349 reported variants but does not cover the full diversity of clinically observed kinase alterations (Saifudeen et al., 2026).
Most importantly, biochemical “hits” require further validation. A drug may inhibit a mutant kinase in vitro but fail clinically because of insufficient exposure, toxicity at required doses, compensatory signaling, lack of tumor dependence on that kinase, or poor activity in cellular and in vivo systems.
Clinical Meaning: A Research Resource With Translational Potential
For OncoDaily Lung readers, the main message is that KIRHub may become a valuable research tool for the next generation of lung cancer precision oncology. It provides a functional map of how clinically approved kinase inhibitors interact with wild-type kinases, oncogenic mutations, and kinase fusions. This can help identify resistance gaps, candidate repurposing strategies, and mutation-specific inhibitor patterns.
The most lung-relevant examples include variant-specific activity across EGFR, ALK, and RET; coverage gaps for MET resistance mutations; and the identification of gilteritinib as a potential inhibitor of selected MET variants in preclinical systems (Saifudeen et al., 2026). These findings need further validation before they can affect patient care, but they provide a strong foundation for translational studies and precision trial design.
In a field where rare variants and resistance mutations increasingly define clinical uncertainty, a systematic kinase inhibitor map can help move drug repurposing from chance observation to rational investigation.
You can read full article here


