The SMART Trial, presented by Pedram Razavi during the ESMO Breast Cancer Annual Congress 2026 , explored how circulating tumor DNA (ctDNA) dynamics combined with therapeutic context may help predict treatment response and disease progression in patients with metastatic breast cancer. Using the personalized tumor-informed Signatera™ assay, investigators developed a clinical support model capable of identifying patients at high risk of progression within 90 days with very high sensitivity and specificity. The findings highlight the growing role of real-time liquid biopsy monitoring in precision oncology.
Background
Managing metastatic breast cancer increasingly depends on understanding whether a patient is benefiting from systemic therapy before clinical or radiographic progression becomes evident. While imaging remains the standard approach for monitoring disease, circulating tumor DNA has emerged as a promising biomarker capable of detecting molecular response and resistance earlier in the disease course.
The SMART platform (Signatera Monitoring for Assessment of Response to Treatment) was designed to integrate ctDNA kinetics with treatment context to estimate the probability of imminent progression in real time. The model aims to provide clinicians with an additional decision-support tool that could potentially guide earlier therapeutic interventions in metastatic breast cancer.
Methods
Investigators evaluated ctDNA in 2,456 patients with metastatic breast cancer using the personalized tumor-informed Signatera™ assay. Treatment decisions and ctDNA testing schedules were determined by treating physicians in routine clinical practice.
The SMART algorithm was trained using ctDNA data from 1,673 patients across multiple breast cancer subtypes and treatment lines:
- Triple-negative breast cancer (TNBC): 368 patients (22%)
- HR+/HER2- disease: 970 patients (58%)
- HR-/HER2+ disease: 101 patients (6%)
- HR+/HER2+ disease: 234 patients (14%)
The model analyzed log-transformed ctDNA kinetic changes during therapy and categorized patients into response or progression groups. A separate test cohort of 783 patients was then used for validation. Progressive disease was inferred through treatment change.
Study Design
SMART analyzed sequential ctDNA pairs to estimate the likelihood of progression within 90 days after testing. The system classified patients into low-risk and high-risk groups for imminent progression.
In the validation cohort, investigators evaluated 3,733 sequential ctDNA pairs. Among these:
- 2,322 ctDNA pairs were classified as low risk
- 1,407 ctDNA pairs were classified as high risk
A total of 590 progression events were identified
Importantly, the model correctly predicted 586 of 590 progression events from high-risk ctDNA pairs, corresponding to:
- Sensitivity: 99.1%
- Specificity: 90.9%
Key Findings
Additional data presented during the congress slides further demonstrated how predicted risk scores correlated with treatment outcomes and time to treatment change.Among 1,360 ctDNA pairs from 348 patients in the validation cohort, increasing predicted risk was associated with progressively shorter time-to-next-treatment (TTNT).
Patients classified as low risk (<20%) had a median TTNT of 9.33 months, while those in the intermediate-risk group (20–80%) had a median TTNT of 3.48 months. Patients categorized as high risk (>80%) had a median TTNT of only 1.97 months, suggesting rapid treatment failure in this subgroup.
The presentation also demonstrated that higher predicted risk scores aligned with higher 90-day event rates across molecular response categories:
- Molecular complete response (mCR) with low risk had a 5% event rate
- Molecular partial response (mPR) with intermediate risk showed a 20% event rate
- Molecular stable disease (mSD) with high risk demonstrated a 69% event rate
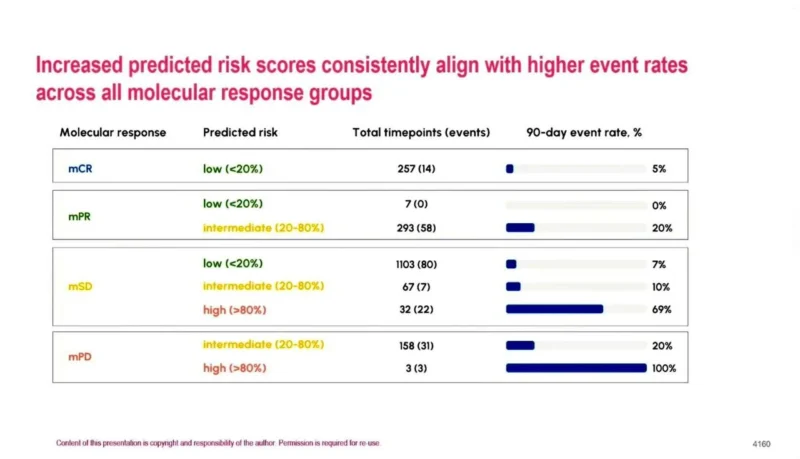
Molecular progressive disease (mPD) with high risk showed a 100% event rate within 90 days
These findings suggest that ctDNA dynamics may provide clinically meaningful information beyond traditional molecular response categories alone.Another important observation involved treatment persistence. In the validation cohort, 98.8% of low-risk ctDNA pairs were not associated with treatment change within 90 days, compared with only 39.7% in the high-risk group. This supports the ability of the SMART model to identify patients likely to experience near-term therapeutic failure.
Conclusion
The SMART ctDNA monitoring platform demonstrated high accuracy for predicting imminent progression in metastatic breast cancer using real-time ctDNA dynamics and therapeutic context. Presented at , the study showed that increasing predicted risk scores were associated with shorter time to treatment change and substantially higher event rates across molecular response groups.


