Divya Nori, Undergraduate Researcher at Eric and Wendy Schmidt Center at Broad Institute of MIT and Harvard, made the following post on X:
“Introducing RNAFlow, a protein-conditioned RNA generative model for structure and sequence design. Excited to present this work, mentored by Wengong Jin, ICML Conference in a few weeks!
Paper.
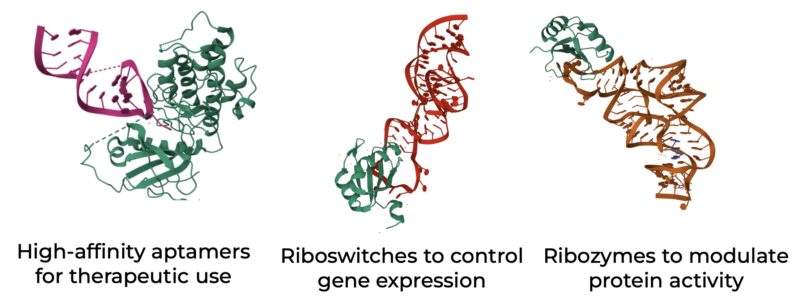
RNAs can be engineered to perform versatile functions, particularly through their interactions with specific protein binding partners.
In the protein design field, diffusion and flow matching models that fine-tune pre-trained structure prediction networks have excelled at conditional binder design. But this is computationally expensive and necessitates a separate sequence design step.
RNAFlow uses inverse folding as a bridge to unlock the generative capability of pre-trained structure prediction models, without fine-tuning. We predict a denoised sequence from a noisy backbone, with protein conditioning (using a modified version of gRNAde from Chaitanya K. Joshi!)
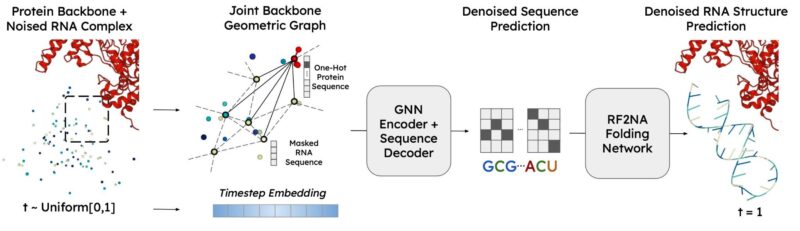
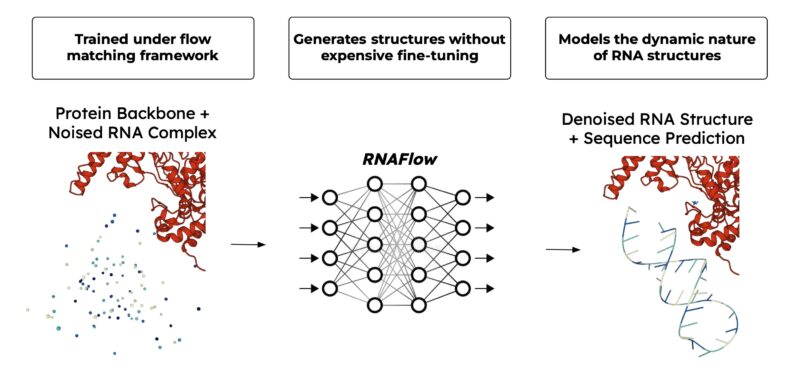
A frozen RosettaFold2NA network folds the sequence into a denoised structure to perform flow matching in structural space. RNAFlow can design RNAs with modest sequence and structure accuracy while maintaining novelty! See the paper for details.
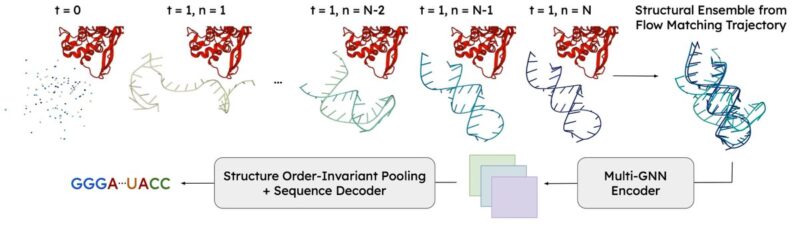
Further, we model the dynamic nature of RNA. Over the course of flow matching inference, RNAFlow generates a trajectory of structures. The final RNA sequence design is conditioned on the last few inference outputs which approximate a conformational ensemble.
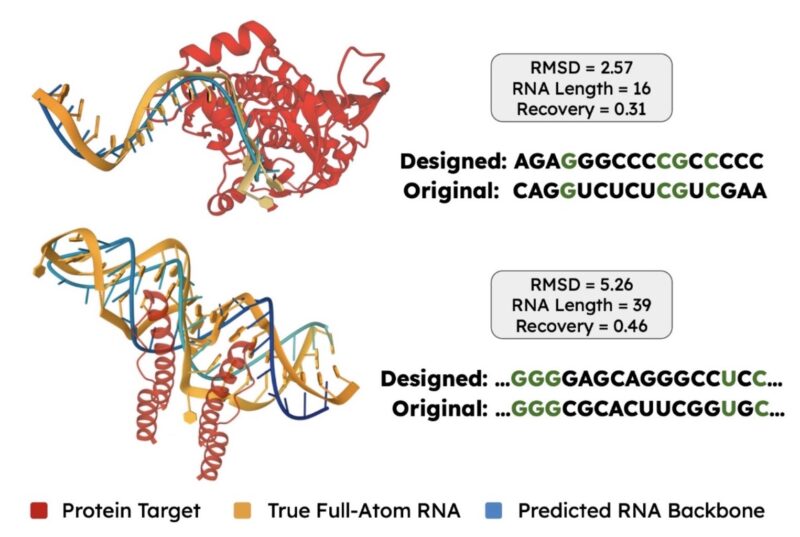
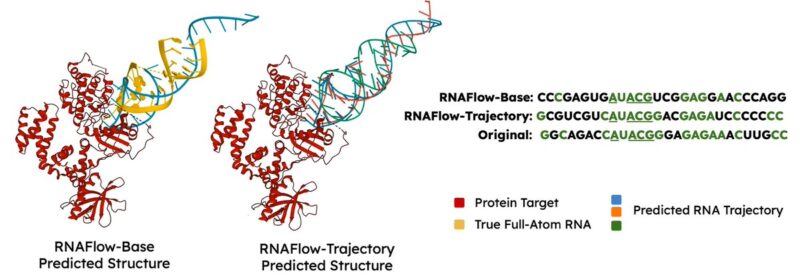
We evaluate whether RNAFlow can design the sequence and structure of an aptamer that binds to GRK2, a target of interest for chronic heart failure. We provide a known binding motif of 4-nucleotides. RNAFlow’s predictions resemble the true aptamer’s sequence and structure!
If you’d like to chat about RNAFlow, or ML for biology in general, please reach out! Excited to connect ICML Conference.”
Source: Divya Nori/X


